Molecular basis of viral neurotropism
Understanding how viruses enter, manipulate, and damage neurons and other cells of the central nervous system.
Virology · Proteomics · Neurotropism · Systems Biology
Postdoctoral Fellow - Stanford University
I am interested in understanding the molecular bases of viral neurotropism. My research combines molecular virology, multi-omics, functional genomics, and computational analysis to understand how neurotropic viruses enter, reprogram, and damage host neurons.

Bio
Alejandro Matía is a CZ Biohub Collaborative Postdoctoral Fellow, working jointly with the Huttenhain lab at Stanford and the Arias group at the CZ Biohub San Francisco. The main focus of his research is to use systems approaches to uncover new insights into viral infections, such as identifying host factors in relevant study models. Alejandro obtained his PhD research at the Spanish National Research Council (CSIC), employing multi-omic technologies such as CRISPR genetic screens and single cell transcriptomics in Poxvirus infections. His interest in Bioinformatics led to the creation of MaGplotR, a tool designed for the analysis of multiple genetic screens. Alejandro also has experience in long-read sequencing, and he has sequenced different viral genomes such as Vaccinia virus, Monkeypox virus, West Nile virus and others.
Research
Understanding how viruses enter, manipulate, and damage neurons and other cells of the central nervous system.
Using proteomics, multi-omics integration, and systems-level approaches to map how viral infection reshapes cellular states and host pathways.
Developing and applying computational approaches to predict how viruses interact with host cells, identify vulnerable pathways, and generate testable hypotheses.
Publications
A preprint describing V-SWITCH, a modular single-vector fluorescent reporter designed to detect live RNA virus infections in living cells.
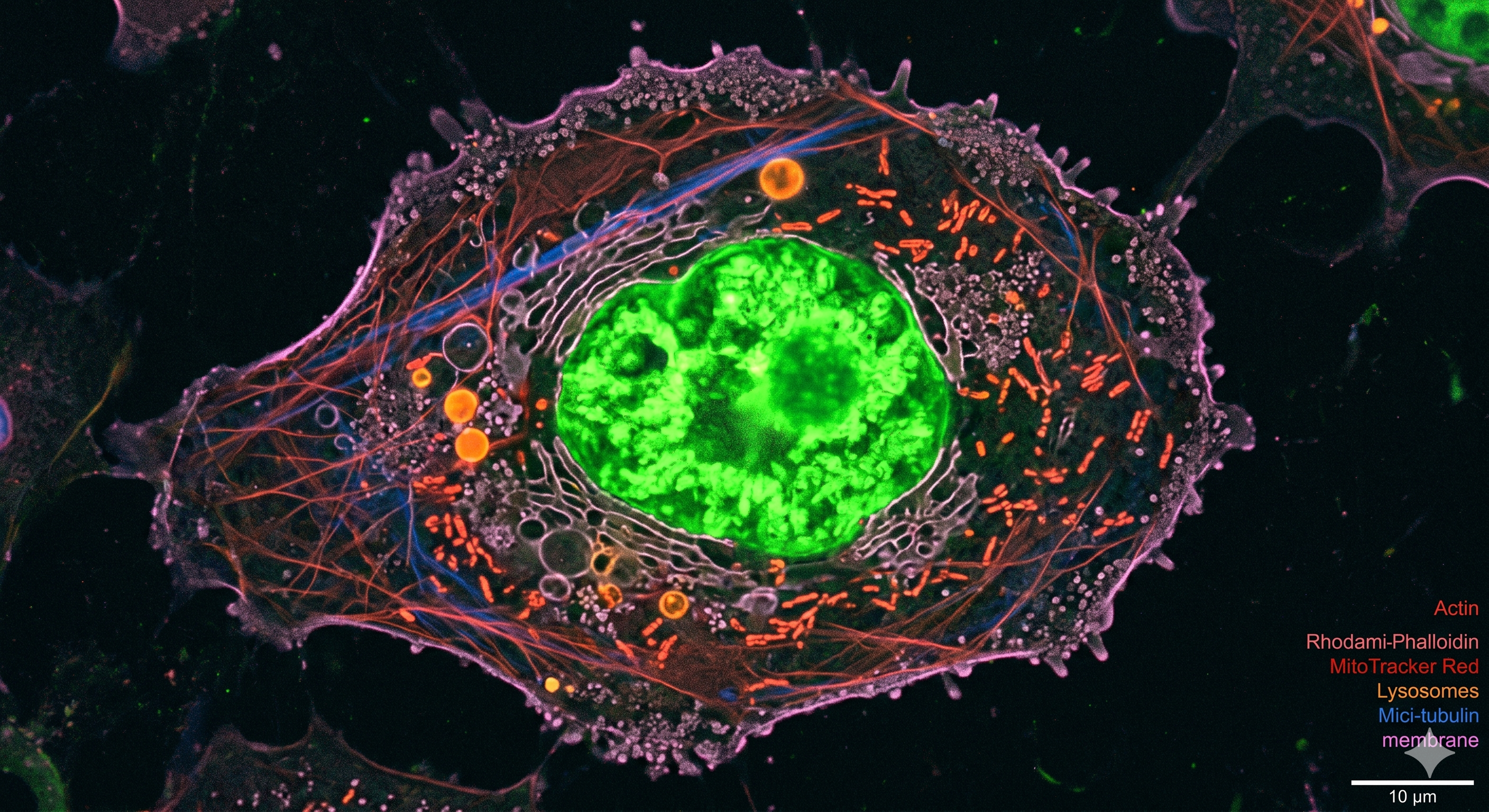
A systems-level study of Vaccinia virus infection dynamics that combines single-cell transcriptomics with proteomics to resolve how host responses evolve across infection.
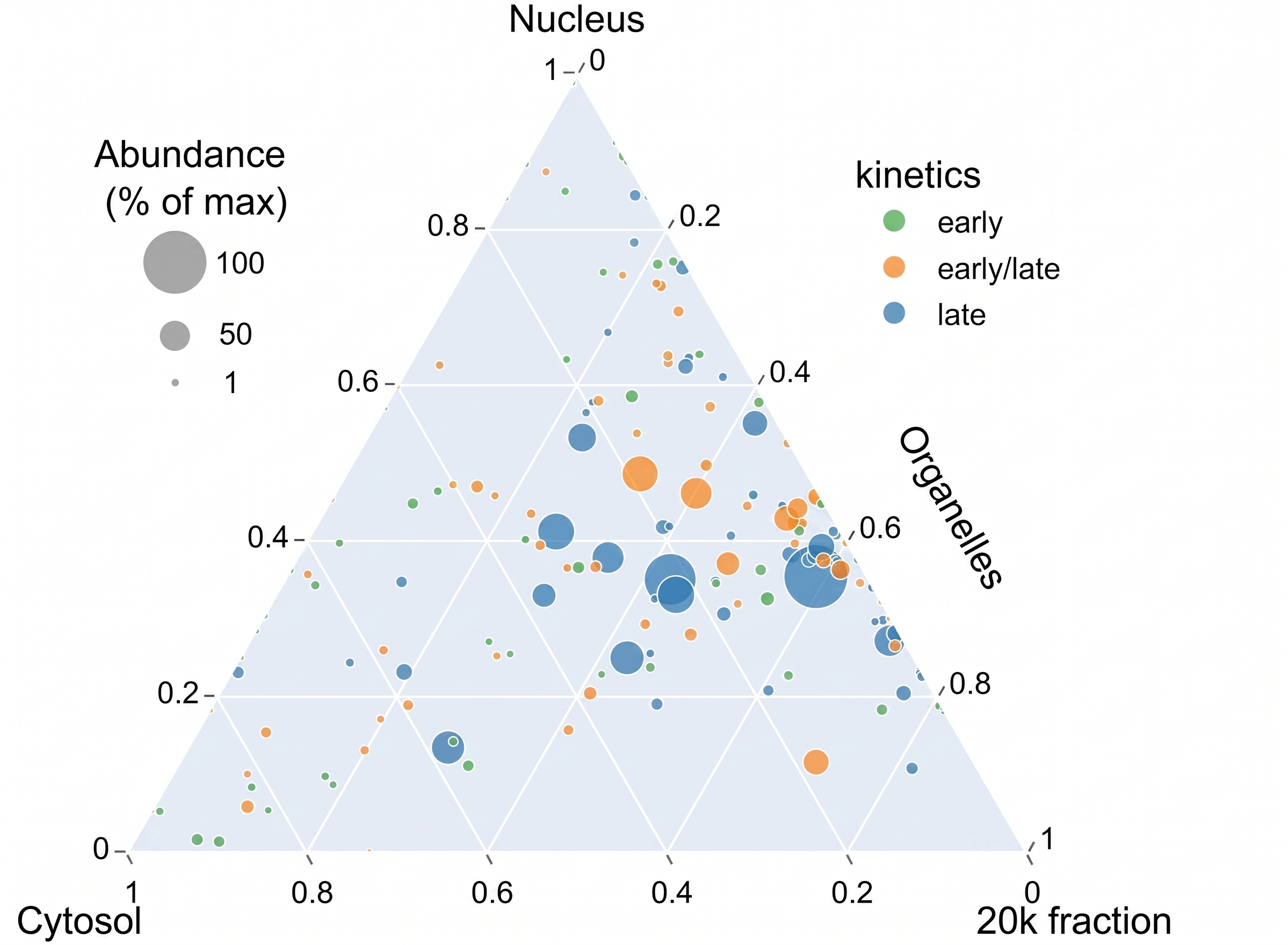
A software-oriented contribution focused on aggregating and visualizing multiple MaGeCK screen datasets in a more streamlined and interpretable way.
Preclinical vaccine study showing robust protection with an MVA platform expressing a prefusion-stabilized SARS-CoV-2 spike, including prevention of detectable brain infection in mice.
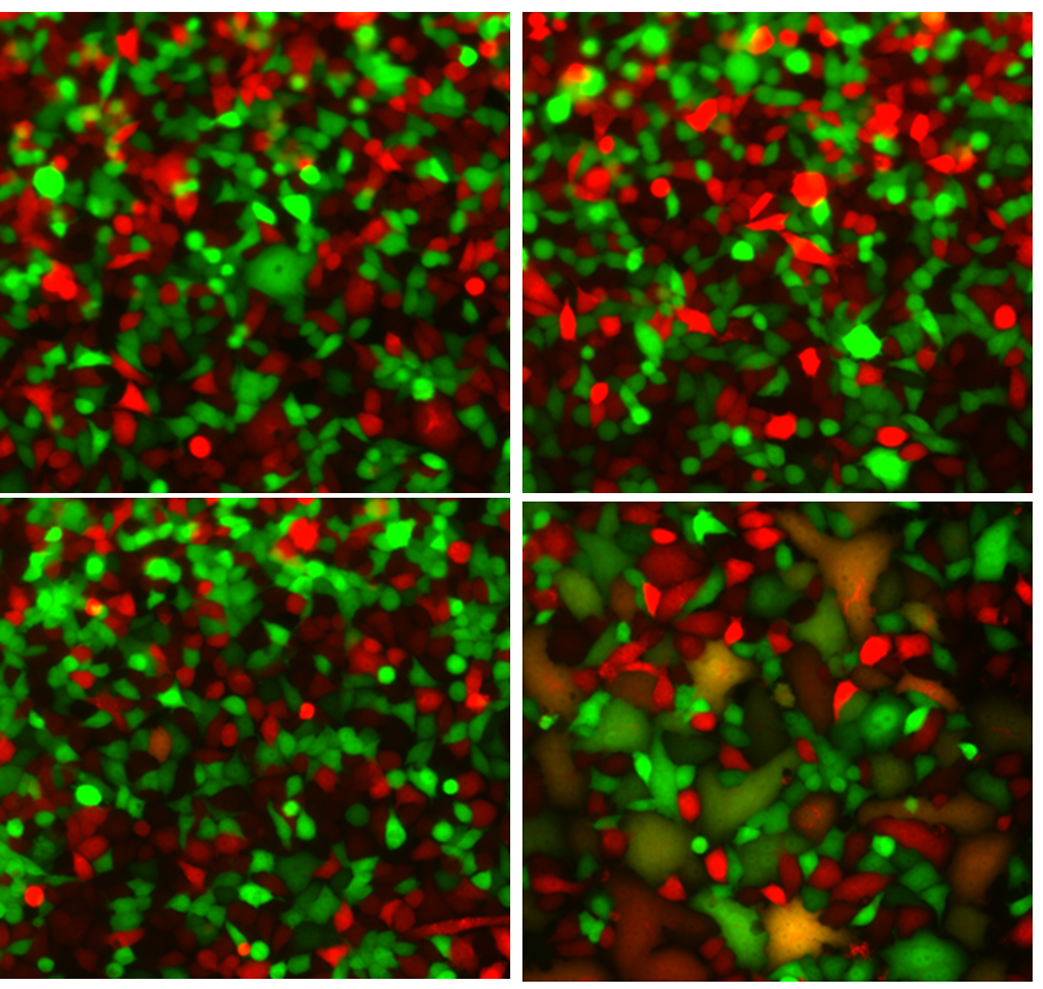
Genome-wide screening work identifying host factors involved in Vaccinia virus infection, including β2-microglobulin as a previously unrecognized entry-related factor.
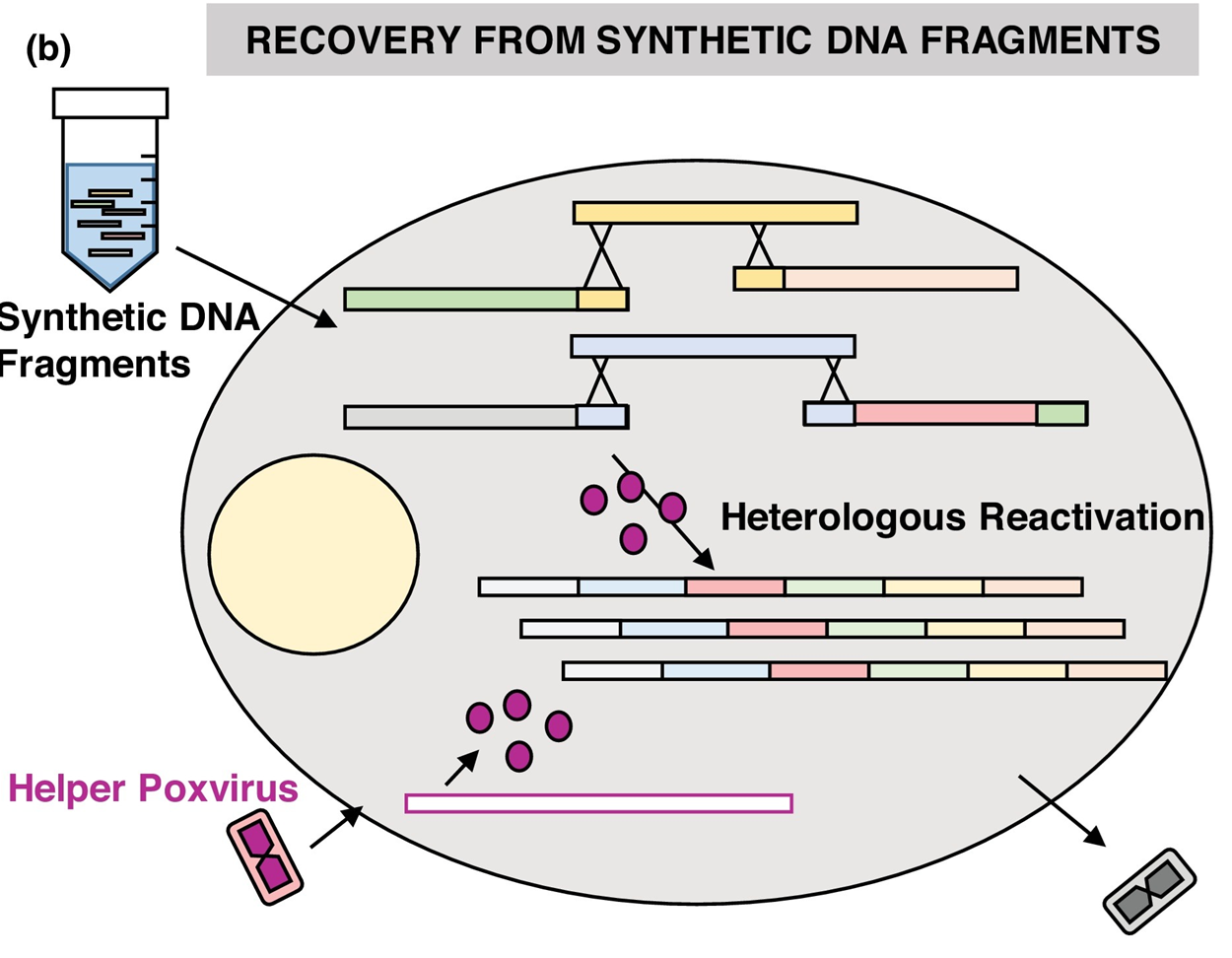
A review of classical and emerging strategies for engineering poxvirus genomes, with relevance to recombinant vaccines, biotechnology, and reverse genetics.
Trajectory
Research Experience
11/2024 – 11/2026
CZ Biohub and Stanford University, CA, USA
11/2023 – 10/2024
Stanford University, CA, USA
07/2019 – 09/2023
INIA-CSIC, Madrid, Spain
09/2019 – 07/2023
INIA-CSIC, Madrid, Spain
07/2022 – 11/2022
Chan Zuckerberg Biohub, San Francisco, CA, USA
01/2018 – 06/2018
Dept. Organic Chemistry, Universidad Autónoma de Madrid, Spain
06/2015 – 06/2016
National Center for Biotechnology – CSIC, Madrid, Spain
Education
09/2019 – 09/2023
Universidad Autónoma de Madrid, Madrid, Spain
06/2021 – 06/2022
Universidad Pablo de Olavide, Sevilla, Spain
09/2017 – 06/2018
Universidad Autónoma de Madrid, Madrid, Spain
09/2012 – 06/2017
Universidad Autónoma de Madrid, Madrid, Spain
Recognition
2026 · Stanford Synthetic Biology Postdoctoral Support — Grant
2026 · ASV Conference Travel Award
2026 · Stanford BioX Travel Award
2024 · Chan Zuckerberg Biohub Collaborative Postdoctoral Fellowship
2019 · Predoctoral Research Fellowship (FPI), Spanish Ministry of Science and Innovation
2018 · Academic Excellence Scholarship, Universidad Autónoma de Madrid
Contact